
 |
|
QGRS Mapper Help: Search & Analysis
The main goal of the QGRS Mapper program is to predict the presence of Quadruplex forming G Rich Sequences (QGRS) in nucleotide entries. QGRS Mapper allows the user to search for putative G-quadruplexes in a variety of ways.
It is possible to enter a nucleotide sequence in raw or FASTA format for analysis. One can search and analyze gene sequences by Gene ID, Gene name or symbol, Accession number or GI number for an NCBI nucleotide sequence entry. The user can opt to change the maximum length of QGRS that will be searched for (the default maximum length being 30) and change the minimum sized G-group tetrads (which is two by default). A screen shot for the Web page for QGRS analysis:
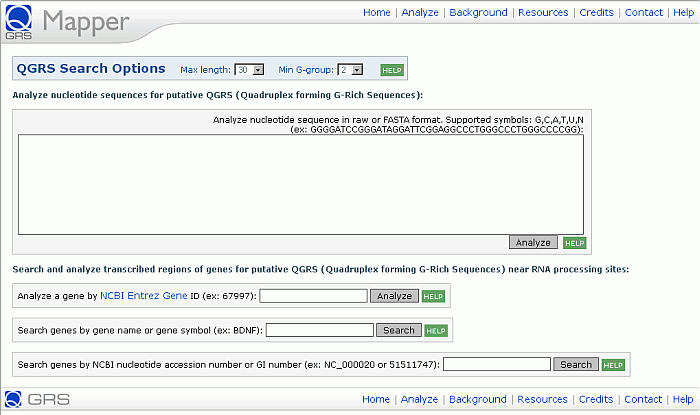
Analysis of User-Supplied Nucleotide Sequences
After entering a sequence in raw or FASTA format, QGRS Mapper will search the sequence for occurrences of QGRS. The user may enter any combination of the letters A, C, T, G, U, N.
Search and Analysis of NCBI Database Entries
There are three ways to connect to NCBI databases for search, retrieval and analysis of gene/nucleotide entries:
1. Gene ID
The Gene ID field allows the user to search the NCBI Entrez Gene database. QGRS Mapper will connect to NCBI, download and parse the gene entry, and then analyze the transcribed region of its nucleotide sequence for the presence of QGRS. For example, entering the gene ID 403437 results in downloading the Brca1 gene sequence for Canis familiaris. QGRS Mapper finds 156 non-overlapping QGRS and 3394 overlapping QGRS in the transcribed region of this gene.
2. Gene Name or Symbol
The Gene Name or Gene Symbol field also allows the user to search the NCBI databases for all such genes. Entering the gene name Bcl2 results in nine different hits which are displayed in this table:

All nine of these entries can be analyzed for the occurrence of QGRS. Clicking on the Gene ID takes the user to the respective Entrez Gene entry. Clicking on the last column initiates analysis of the selection by QGRS Mapper.
3. Accession Number or GI Number
Similarly, the user can also enter an NCBI accession number to search for gene sequences. For example, searching the accession number AF312033 results in 12 hits being displayed for this RefSeq nucleotide sequence entry. In the table below, The Gene Symbol field links to the corresponding NCBI GenBank entry. The Number of Products field refers to the number of alternatively processed mRNA products.
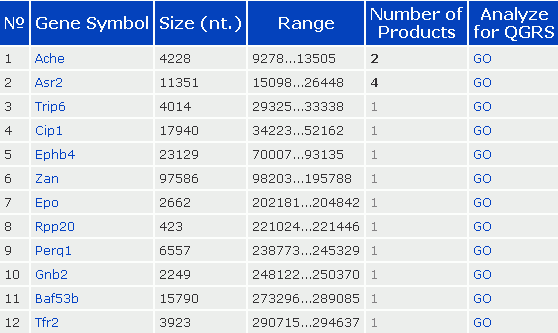
The search phase of the program is followed by an analysis of the QGRS contained in the query sequence. In this phase of QGRS Mapper, the sequence data previously downloaded is analyzed to identify and map all QGRS relative to locations such as splice sites in exons/introns, and poly-A site (if these locations are known). Furthermore the QGRS are scored. The computed G-score is used to eliminate overlapping QGRS.
At times, QGRS Mapper must analyze a considerable amount of data. For example, the mouse version of the gene PTPRU, which is 69822 bases long, contains 94681 QGRS of length up to 45 bases. QGRS Mapper will find, analyze, and map all of these sequences. During this analysis a message is displayed indicating the estimated time left to completion.
QGRS Mapper Output
After the analysis of overlaps is completed, QGRS Mapper displays a summary of its findings, in the Gene View. This summary includes basic gene information such as the gene ID, gene symbol, gene name, a link to the NCBI entry, organism name, chromosome number, and number of products and polyA signals. Information is also given for each product, such as the number of exons and introns, number of QGRS (non-overlapping and overlapping), number of QGRS found near RNA processing sites, and a visual map of each RNA product.
As an example, the Gene View for the GREB1 gene is displayed in following figure, showing the table of gene information and product information.

At this stage in the analysis the user can choose among three further displays: "Data View", "Data View (with overlaps)", and "Graphics View". This can be done for the entire gene or for any particular product.
In the Data View, a table is displayed showing information for each of the set of non-overlapping QGRS. This table displays the position of the QGRS, which exon/intron it appears in, its distance from 3' and 5' splice sites, the QGRS sequence (with each G-group underlined) and the corresponding G-score. Similar display is also shown for each QGRS mapped to polyA region in the product. If the user requests the Data View for the entire gene, then the QGRS information is shown for each product. The "Data View (with overlaps)" gives the same information but shows the locations of all QGRS. The following figure shows fragment of the Data View for product 1 of the GREB1 gene:

The user can also choose the Graphics View to give a visual display of the location of QGRS. This allows the user to see the location of QGRS relative to exons and introns (if that information is available). The Graphics View has the following components:

The "Graphics" View for the entire GREB1 gene may be seen in this figure:

 |
|
Visitors 

| |